|
However, since coronaviruses have many mutants, new primers specific to the target strain must be continuously developed for sequencing. Currently, a method for amplifying a genome fragment using a strain-specific primer set is being used ( Geng et al., 2012 Zhang et al., 2020). In 2004, the complete genome of HCoV-NL63 was sequenced using the Virus-Discovery-cDNA AFLP (VIDISCA) method ( van der Hoek et al., 2005). The first genome of HCoV-229E, reported in 2001, was sequenced by synthesizing it as cDNA and cloning it into the vaccinia virus ( Thiel et al., 2001). To solve this problem, various alphacoronavirus genome sequencing methods have been developed over the past 20 years. Since the viral load in clinical specimens is minimal, it is difficult to directly recover a sufficient amount of nucleic acid for genome sequencing. The use of whole genome sequencing to identify SARS-CoV-2 mutants is a good example ( Wang et al., 2020 Zhou et al., 2020 Hu et al., 2021). Thus, whole-genome sequencing is essential to determine whether a virus is a mutant strain ( Gilchrist et al., 2015 World Health Organization, 2020). Moreover, any viral variant located beside the short marker gene region cannot be determined through the conventional standard rRT-PCR. However, false-negative results may occur when there is a mutation in the marker gene region ( Kevadiya et al., 2021). ORF1a and b ( van der Hoek et al., 2005), and nucleocapsid genes ( Bergner et al., 2020 Morehouse et al., 2020) are used as diagnostic markers. 
These two types of alphacoronaviruses are known to cause 30% of common colds in humans ( Su et al., 2016).Īlphacoronaviruses are diagnosed using real-time reverse transcription-polymerase chain reaction (rRT-PCR). Two types of alphacoronaviruses infect humans: HCoV-229E, which was first described in the mid-1960s, and HCoV-NL63, which was identified in 2004. Unlike betacoronaviruses, which include deadly species such as MERS-CoV and SARS-CoV, alphacoronaviruses have received relatively little attention due to their low fatality rate ( Vassilara et al., 2018). Both alpha- and beta- coronaviruses are known to cause diseases in humans. They are classified into four distinct genera: alpha, beta, gamma, and delta. The AC primer set and experimental procedure optimized in this study may enable the fast diagnosis of mutant alphacoronaviruses in future epidemics.Ĭoronaviruses have had a major impact on mankind since the mid-1960s. In the mock clinical specimen, the detection limit was 10 −3 PFU/ml (10 2 copies/ml). The sequencing accuracy varied depending on the sequencing technology, but all sequencing methods showed a sequencing error of less than 0.01%. The amplicon library sequencing data were assembled into complete genome sequences in all sequencing strategies examined in this study. After genome amplification, an evaluation using various sequencing platforms was carried out. The selected primers, named the AC primer set, were composed of 10 primer pairs stretching over the entire genome of alphacoronaviruses, and produced PCR products of the expected size (3–5 kb) from both the HCoV-229E and HCoV-NL63 strains. To this aim, we designed overlapping primer pairs capable of amplifying the entire genome of all known human alphacoronaviruses. Thus, in this study, we aimed to develop a universal primer set valid for all human alphacoronaviruses and applicable to samples containing trace amounts of the virus. At present, there are no universal primers available for alphacoronaviruses and that, since these viruses have diverse strains, new primers specific to the target strain must be continuously developed for sequencing. 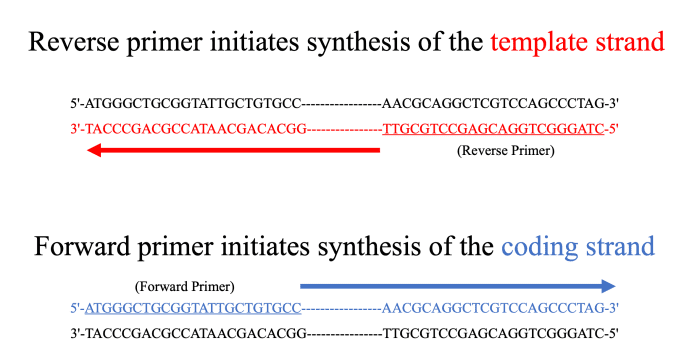
However, due to the low viral titers in clinical samples, certain amplification steps are required for viral genome sequencing. Rapid and accurate sequencing covering the entire genome is essential to identify genetic variations of viral pathogens.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
